Multiomics data analysis parameters¶
The Multiomics configuration file allows you to integrate different data types and, as a default we integrate clinical and proteomics data. In this context, the configuration file includes a multi-correlation section where all clinical and proteomics data are correlated and depicted as a network. Another option is to run a Weighted Gene Co-expression Network Analysis (WGCNA), where features (proteins) are clustered in co-expression modules and further correlated to the clinical variables.
The multiomics default analysis pipeline can be accessed here.
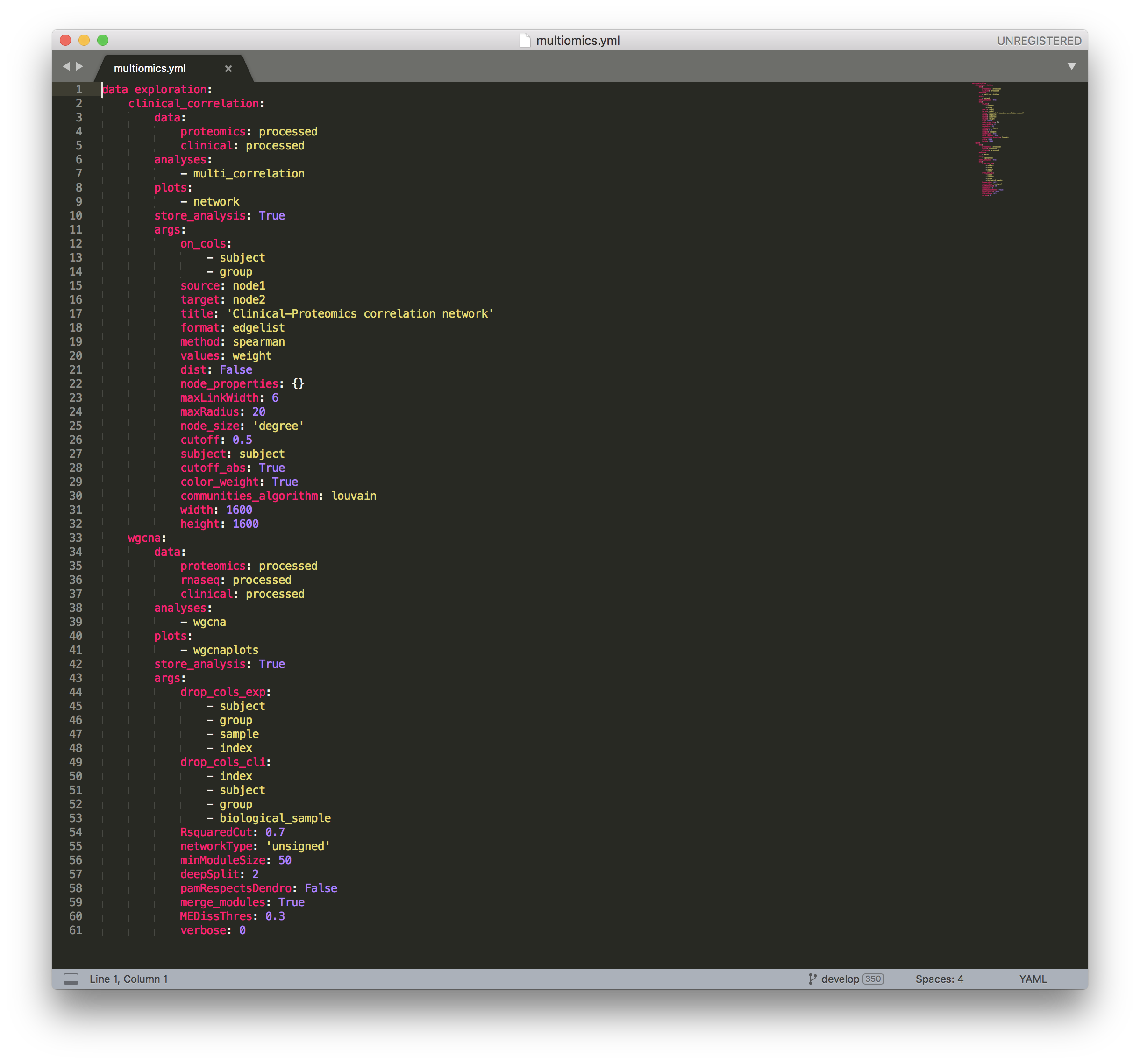
Multiomics configuration file¶